Michael M. Zeineh, *Paul M. Thompson, Stephen A. Engel, Susan Y. Bookheimer
Brain Mapping Division and *Laboratory of Neuro-Imaging, UCLA School of Medicine, Los Angeles, CA USA
ABSTRACT
Introduction
The medial temporal lobe plays a vital role in memory, but activation has been elusive and difficult to characterize in neuroimaging studies of memory. In particular, such studies often have to compromise either resolution or statistical power. Especially in event-related experiments, neuroimaging studies often scan multiple subjects and employ group statistics to generate robust maps of activity. However, such methods generally sacrifice spatial resolution in order to compensate for anatomic variability; in doing so they lose the ability to specifically define the locus of activation in the medial temporal lobe. With single-subject data, one can achieve high spatial resolution, but in comparison with group analyses, this lacks both statistical power and the ability to visualize activity representative of the subject pool. Here we present a new method that maintains high resolution while combining data across subjects. Our previous work involved generating flat maps of the medial temporal lobe, which allow us to visualize activity in the various substructures of the region (1,2). By further using elastic warping techniques, flat maps of multiple subjects can all be transformed into the same space, where powerful group statistics can then be performed. We demonstrate its use in a novelty-encoding experiment.
Methods
Eight subjects underwent scanning on a 3T GE scanner with ANMR upgrade for EPI. The novelty paradigm consisted of 5 repetitions of 16 novel pictures followed by 16 repeated pictures (ISI 2.55 sec). Unfolding methods were identical to (1,2). High-resolution structurals (512x512, 3mm thick) coplanar with high-resolution functionals (128x128) were acquired. We segmented, unfolded, and demarcated the boundaries of the hippocampal subregions using the structural images. Motion-corrected functional images were mapped onto the structurals to produce flattened timeseries images. Single-subject activation maps were generated by smoothing the timeseries (4mm FWHM), and correlating signal intensity with a smoothed boxcar function.
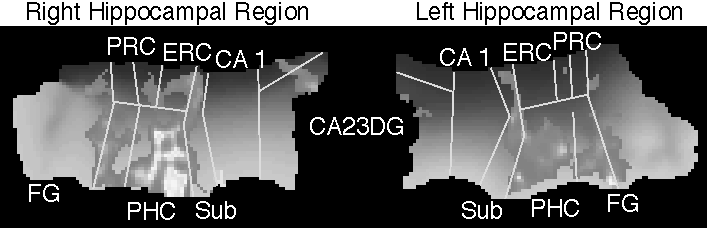
To register subjects in the same space, we manually outlined the demarcated boundaries between the hippocampal subregions (CA23DG, CA1, Sub, ERC, PRC, PHC, FG). An elastic warping technique based on continuum-mechanics (3) generated transformations that map from one hippocampus to another by smoothly overlaying the boundaries. Applying this transformation to correlation maps yielded 8 coregistered correlation maps for each hemisphere. For each flat voxel, the mean and the variance of the subjects’ correlation values was used to generate a t-map that was thresholded at 3.0.
Results
Consistent with published results, the statistical maps show clear activation of the parahippocampal cortex and fusiform gyrus. Activation further included the bilateral subiculum and anterior CA23DG, while on the right there was further entorhinal and perirhinal activation. No CA 1 field increases were present.
Discussion
The combination of unfolding methods with warping techniques delivers higher resolution than that offered by traditional warping-based volumetric methods. Segmenting the gray matter manifold allows the use of 2-D rather than 3-D smoothing, preserving the distinction between regions adjacent in 3-D space but disparate on the hippocampal surface. The unique anatomy of the hippocampal region further facilitates the standardization of boundary definitions, which we have utilized to cross-register subjects into the same space. By developing a standardized template to register hippocampal activations, we will ultimately be able to integrate the current diverse array of memory findings.
References
1.Zeineh et al., (1998). HBM 1998, S693. 2. Zeineh et al., (1999). HBM 1999, S982. 3. Thompson et al., (1996/7). Medical Image Analysis, 1(4):271-94.
Related Publications
(back to main list)
Contact Information
Paul Thompson
73-360 Brain Research Institute
UCLA Medical Center
10833 Le Conte Avenue
Westwood, Los Angeles CA 90095-1761, USA.
| RESUME| E-MAIL ME| PERSONAL HOMEPAGE| PROJECTS |
|---|